This story is part of The Cancer Letter’s ongoing coverage of COVID-19’s impact on oncology. A full list of our coverage, as well as the latest meeting cancellations, is available here.
NCI’s Frederick National Laboratory for Cancer Research has launched three initiatives focused on SARS-CoV-2:
Identifying genetic determinants of SARS-CoV-2 susceptibility and outcomes at the Cancer Genomics Research Laboratory,
Testing and validating serologic assays for SARS-CoV-2 in the Serology laboratory of the Vaccine, Immunity, and Cancer Program, and
High-throughput screening for small molecule inhibitors of SARS-CoV-2 proteins, with technology developed by the RAS Initiative.
“We think that it was built for a situation like this, where speed, flexibility, and expertise are critical to addressing such a deadly public health threat,” Douglas Lowy, NCI principal deputy director, said April 9 in an emergency virtual meeting of the NCI Board of Scientific Advisors and the National Cancer Advisory Board.
“Of course, what we are doing at the Frederick Lab—even more broadly across NCI—is only part of a truly massive global effort to understand, control, and overcome the pandemic.
“No one research group, institution or country can do this alone, but because of what we’re trying to do, I like to think that we are trying to take a massive mountain, an infectious threat as large as Mount Everest, and, ultimately, be able to subdue it into a molehill that, with vigilance, can be restrained and tamed.”
The National Institute of Allergy and Infectious Diseases has used the Frederick National Lab to respond to other epidemics, including SARS in 2003, Ebola in 2013, and Zika in 2015. NIAID is sponsoring a trial of the antiviral agent remdesivir, which was originally developed for treatment of Ebola and Marburg virus infections.
Remdesivir, a nucleoside analog that functions as an RNA chain terminator, was subsequently found to inhibit replication of other RNA viruses, including coronaviruses. The drug, made by Gilead Sciences, is the focus of multiple trials worldwide and is closely watched by the public health and financial communities.
No one research group, institution or country can do this alone, but because of what we’re trying to do, I like to think that we are trying to take a massive mountain, an infectious threat as large as Mount Everest, and, ultimately, be able to subdue it into a molehill that, with vigilance, can be restrained and tamed.
Earlier this week, The New England Journal of Medicine published a paper on remdesivir. At this writing, the market is moving in part as a result of a news report based on fragmentary information from the University of Chicago cohort in two Gilead phase III studies.
As data accumulate, Gilead is scaling up production of the drug, with a target of more than 1 million treatment courses by December.
“We have proactively and rapidly scaled our supply chain,” the company said in a recent statement. “As of late March, using the active ingredient we already had in our inventory, we have increased our supply to more than 30,000 patient courses of remdesivir on hand, assuming a 10-day course of treatment for patients. As new raw materials arrive over the next few weeks from manufacturing partners around the world, our available supply will begin to rapidly increase.
“Every day we are improving processes, shortening timelines and increasing volumes as we work to bring remdesivir to patients as soon as possible. Our goal is to produce a total of:
More than 140,000 treatment courses by the end of May,
More than 500,000 treatment courses by October,
More than 1 million treatment courses by December,
Several million treatment courses in 2021, if required.”
The NIAID global therapeutic trial of remdesivir in COVID-19 patients has, to date, enrolled over 500 patients in eight countries: Denmark, South Korea, Germany, Singapore, Greece, the United Kingdom, Japan, and in the United States, Lowy said at the BSA and NCAB meeting.
The Adaptive COVID-19 Treatment Trial is scheduled to be completed on April 1, 2023. While it appears the trial has exceeded its accrual goal of 440 patients, it’s unclear whether enrollment would be defined as completed, as per protocol. Preliminary results are expected later this spring.
Lowy’s remarks to BSA and NCAB follow:
Ned [Sharpless, NCI director], thanks for reviewing some of the responses that we have made at NCI and the extraordinary efforts extramurally during this really terrible time.
I want to really tell people that Ned has not only been working 24 hours a day, so we know that he’s been working hard. But let me assure you, he has also been working smart and he is everywhere trying to, essentially, deal simultaneously with the ramifications of the COVID-19 epidemic, as well as being able to continue cancer research.
I am going to be focused on three different projects that are being conducted largely at the Frederick National Laboratory, because I think that they exemplify our ability there to change things pretty rapidly. For those of you who may not be entirely familiar with the Frederick National Laboratory, it was established in 1971 by the National Cancer Act and it’s the only one of the 42 Federally Funded Research and Development Centers in the United States that is dedicated to biomedical research.
The Frederick National Lab is sponsored by the NCI, but it’s operated by a private contracting firm, Leidos Biomedical Research, and Ethan Dmitrovsky, whom many of you know from his time at Dartmouth and then at MD Anderson. He’s the head of the Frederick National Laboratory. So, a lot of cancer research is performed at the Frederick National Lab and includes lots of different things, from large scale genomic and proteomic studies, to advanced biomedical computing research.
The lab also anchors many NCI initiatives, such as the RAS Initiative and the Cryo-Electron Microscopy Facility. I’m not going to talk specifically about the Cryo-EM Facility, but that facility, which is for the exclusive use of the extramural community, is being made available for investigators who want to solve Cryo-EM problems related to the COVID-19 epidemic.
The Frederick National Laboratory houses two current Good Manufacturing Practice facilities that produce experimental treatments for first-in-human clinical trials.
One is for NCI and the other is for NIAID, and NIAID and NCI are the major users of Frederick. Let me turn briefly to a discussion of NIAID. They have helped to support and lead clinical trials in two therapies that proved to be particularly effective for treating Ebola infections, including a monoclonal antibody that, actually, was developed and manufactured by NIAID’s vaccine research center through the Frederick National Laboratory.
With the current epidemic, they have stood up a clinical trial of remdesivir, and for those of you who don’t know about it, it’s a nucleoside analog, which is an antiviral that functions as an RNA chain terminator.
It was originally developed for treatment of Ebola and Marburg virus infections, but it was subsequently found to inhibit replication of other RNA viruses including coronaviruses. The next slide shows you the patient accrual for the remdesivir trial sponsored by NIAID, but stood up by the Frederick National Laboratory.


The request came from NIAID at the end of January and the trial was started at the end of February. And, as of a few days ago, there were more than 500 patients enrolled in the trial.
The majority of the patients come from the United States, but they are scattered throughout other parts of the world, patients who also are enrolled in this international trial.
So, having heard a bit about NIAID, let me tell you about three different projects which the NCI has begun largely, although not exclusively, at the Frederick National Laboratory. And I think about this as pivoting some cancer research activities at Frederick to the SARS-CoV-2 research.
Genetic determinants
The first project that I’ll mention is identifying genetic determinants of SARS-CoV-2 susceptibility and outcomes. The second is testing and validating serologic assays for SARS-CoV-2, and the third is high-throughput screening for small molecule inhibitors of SARS-CoV-2 proteins.
Let me go through them one at a time.
We’re going to talk about identifying genetic determinants of susceptibility and outcomes. This is under the auspices of the Cancer Genomic Research Laboratory headed up by Stephen Chanock and largely staffed by people who are from the Frederick National Laboratory.
The goals of this project is to rapidly identify variants which could identify lead to targets for therapy, insights into the biology of COVID-19 pathogenesis, and use for screening and public health. And it really will be using genome-wide association studies, or GWAS studies, looking for single nucleotide polymorphisms or mutations. A very important principle of everything that is done at the Frederick National Laboratory related to the COVID-19 epidemic is that the information that data will immediately be shared with the community.
The next slide tells you about really two things. The first is that the COVID-19 epidemic probably has much more in common with HIV, where you have a single etiologic agent than with cancer, which is such a complicated situation, and where GWAS analyses have not yielded quite the same degree of insight that have been seen with HIV.


The second reason is, that Mary Carrington did pathbreaking HIV research—and she is part of the Frederick National Laboratory, but also affiliated with the Center for Cancer Research—and this slide is taken from a recent review, where she talks about the different mutations that there are with CCR5, which is the main receptor by which HIV binds to T-cells and gets into them.
Her laboratory identified the so-called delta 32 mutation, which greatly reduces your risk of infection. But the research related to CCR5 has identified mutations that decrease risk and increase risk of infection and rate of disease progression, and, ultimately, has also led to the development of inhibitors for CCR5 interaction that have been FDA approved.
I now just want to tell you very briefly about the three cohorts that are going to be followed.
One is an Italian epidemic cohort, this is in collaboration with NIAID, and it’s likely to be skewed to patients who have a poor outcome, because of how poorly so many people have done in Italy.
A second project is through the NIH Clinical Center, and this is in collaboration with the Genome Research Institute as well as NIAID, and it is centered at the moment at the clinical center, but it is planned to be able to be expanded to extramural centers. And when that expansion is available, we hope to go out to different centers and make people aware of it.
The third is a longitudinal cancer cohort, and Jim Doroshow is going to discuss that in some detail. But this cohort will be looking at infections focused on cancer patients, which, of course, are a group that is of increased risk for poor outcomes. But we expect that the cohort will include patients with a benign course, in addition to those with poor outcomes, and there will be detailed prospective information.
And we hope that it will be possible to identify through the genome-wide association studies mutations that are associated with increased risk of poor outcome and others of a decreased risk of a poor outcome.
Serologic assays
The second project that I want to discuss is the serology assay. And before I get to that, I want to show you that this is really a highly specialized program, which has been used up to now primarily for HPV serology. And it is headed up by Ligia Pinto, who’s the director of the Vaccine, Immunity and Cancer Program.
Ligia’s serology laboratory is multifaceted and is involved in collaborations with extramural HPV vaccine community, supporting vaccine trials sponsored primarily by NCI, and, also, is the HPV Serology Standardization Initiative for the World Health Organization and it is jointly supported with The Bill and Melinda Gates Foundation.


This shows you data from Ligia’s serology laboratory, which was published six weeks ago from the Costa Rica Vaccine Trial. These are post hoc data, but they show that, over an 11-year period, that the young women who were immunized with just one dose of the GlaxoSmithKline HPV vaccine, their antibody levels to HPV-16 remained high throughout this 11-year period.
And I should also point out again, post hoc analysis, all of these women, none of them have been infected with HPV-16, in contrast to the control. And this has led to a large clinical trial in Costa Rica to test the hypothesis that one dose of the vaccine will be able to provide strong long-term protection against the HPV types targeted by the vaccines.
So, the notion is to convert part of the HPV serology lab to focus on SARS-CoV-2 serology. And this is a collaborative research effort with multiple laboratories, NIAID, CDC, Mount Sinai, and other academic laboratories—and the interaction, cooperation has simply been phenomenal.


The short term goals are to characterize performance of different serological assays and to correlate them with neutralization assays, so we can also understand possible cross reacting sera from prior to the epidemic. This will be important because, as most of you are aware, the notion of getting people back to work, a particularly attractive group would be those people who are antibody-positive, and so, it’s critically important to make sure that the people who are antibody positive, these are not false positives.
Because of the interactions that we are having with various parts of the U.S. government, as well as amazing cooperation through NCI cancer centers and Ned’s initiative, the FDA has asked us if we would essentially help to validate the serological tests that have been submitted to the FDA. That is in progress, but I want to reiterate, it’s being done as a wide collaborative in Denver.
Longer-term goals will be to understand the implications of being seropositive. We think that people are going to be resistant to reinfection, but we don’t know for a fact that that is the case, and this will be a critically important issue.
Another issue is the duration of seropositivity. And then, longer-term, to participate just for the serology effort in some of the longitudinal efforts, particularly for the cancer patient effort to be discussed by Dr. Doroshow.
Small molecule inhibitors
The third and last topic is the high-throughput screening for small-molecule inhibitors and the technology, actually, was developed by the RAS Initiative. So, Kevan Shokat and his colleagues at UC San Francisco, back in 2013, essentially recognized that you could attach, covalently, to cysteines to make subhydro bonds with small-molecule inhibitors. And his laboratory focused on RAS glycine-12 cysteine mutant alleles.
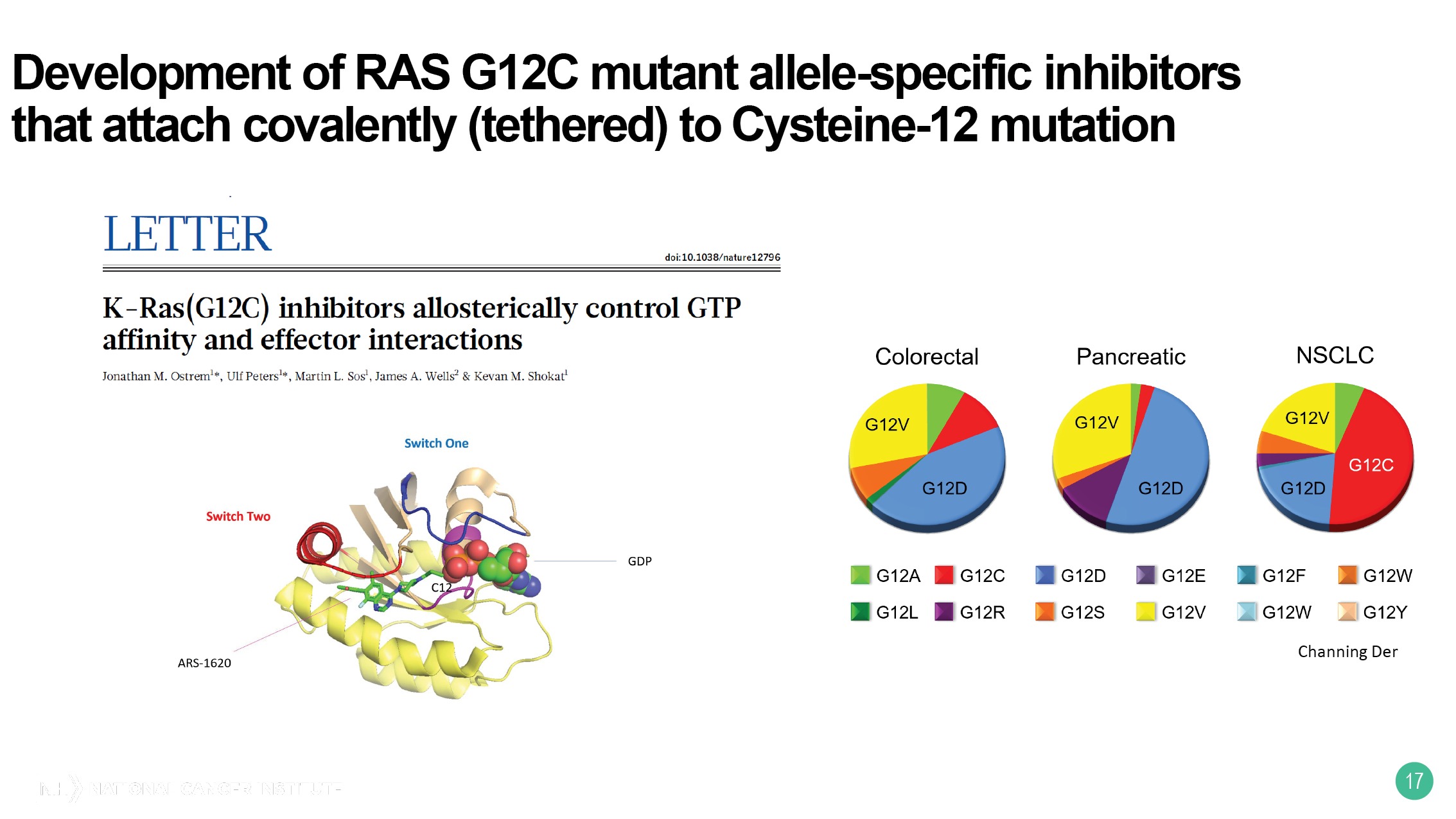

This is a relatively common mutant allele in non-small cell lung cancer, but uncommon unfortunately, in this context in colorectal cancer and pancreatic cancer. The next slide shows you that a number of different pharmaceutical companies have taken this idea and developed inhibitors against G12C.
Early phase trials have been completed, they have been quite encouraging, and phase II and ultimately phase III trials are going to be ongoing. And we hope that this will end up leading to improved outcomes, at least for the subset of patients with mutant KRAS who have the G12C mutation.


So, this approach can be used not just for G12C, but actually for any situation where there is a pocket that might be targeted with a small inhibitory molecule, providing that there is a cysteine adjacent to that pocket. And in the figure it shows the cysteine and then under reducing conditions, you get the subhydro groups connecting to each other, forming the covalent bond and the inhibitor bound to this.


For the FNL disulfide tethering library, David Turner at the Frederick National Laboratory in the RAS Initiative has been the major person to take essentially 13,000 of carboxylic acid building blocks and essentially to screen them, so that you screen out undesirable characteristics and screen in more desirable characteristics.
And reduce this, then, to a library that’s about 10% that of the original library, about 1,200 unique disulfide fragments. And what’s shown with a hierarchic clustering is that this library is quite diverse.


So, colleagues at the Argonne National Laboratory, the Department of Energy in Chicago have been essentially identifying with the two proteases encoded by the coronavirus that they have adjacent exposed cysteine residues to the areas where there might be a binding pocket for an inhibitor. And so, the Frederick National Laboratory is going to take its tethering library and screen the tethering library for such possible inhibitors.


So, once the lead compounds are identified, medicinal chemists at the University of Chicago are going to essentially be optimizing this, going to, also at the Argonne National Laboratory, to use artificial intelligence for further optimization. And then, it will be in vitro inhibition against the proteases, and, ultimately, interference with infection by the authentic coronavirus.


So, I’ve tried to give you a smattering of what is being done at the Frederick National Laboratory, and we think that it was built for a situation like this, where speed, flexibility, and expertise are critical to addressing such a deadly public health threat.
Of course, what we are doing at the Frederick Lab—even more broadly across NCI—is only part of a truly massive global effort to understand, control, and overcome the pandemic. No one research group, institution or country can do this alone, but because of what we’re trying to do, I like to think that we are trying to take a massive mountain, an infectious threat as large as Mount Everest, and, ultimately, be able to subdue it into a molehill that, with vigilance, can be restrained and tamed.
Our patients are counting on us, and we must not let them down.













